Group Haselbach
David Haselbach
Research Institute of Molecular Pathology (IMP), Campus Vienna Biocenter 1, 1030 Vienna, Austria
Phone: +43 (0)1 79730-3009, Email: david.haselbach(at)imp.ac.at
Watching Molecular Machines in action
Our lab studies the design principles of macromolecular machines that enable them to translate the thermal noise power of the environment into productive conformational changes. We are combining classical biochemical and biophysical methods with cutting edge electron microcopy to find out what are the nuts and bolts that make the machines work. Our central aim is thereby to reconstruct trajectories of molecular machines in action in the highest possible spacial and temporal resolution.
Key Publications
* Haselbach D, Komarov I, Agafonov DE, Hartmuth K, Graf B, Dybkov O, Urlaub H, Kastner B, Luehrmann R, Stark H (2018). Structure and Conformational Dynamics of the Human Spliceosomal Bact Complex. Cell 172, 1–11, doi: 10.1016/j.cell.2018.01.010
* Haselbach D, Schrader J, Lambrecht F, Henneberg F, Chari A, Stark H (2017). Long-range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs. Nat Commun; 8:15578. doi: 10.1038/ncomms15578.
Projects within VBC Ubiquitin Club
Despite their intricate design, molecular machines can fail. Using the cryo electron microscopy, we are trying to understand the molecular response of molecular machines on stress and disease. In our initial study, we focused on the molecular adaption of the human 26S proteasome towards chemical inhibitors. Proteasomal degradation is a multi-step process involving substrate recognition, de-ubiquitination and unfolding carried out by the regulatory particle and proteolysis performed by the core structure. Even though it has been well documented that the proteolytic activity of the core is shut down by using covalent inhibitors, the effect on the holoenzyme was unknown.
We found a major allosteric effect that translates the binding of a single inhibitor to the proteolytic site to a restriction of the conformational space of the regulatory particle that is more than 15 nanometres away. We hypothesise that this can be sensed by different rescue factors that bind to the proteasome, resulting in a release of the applied stress.
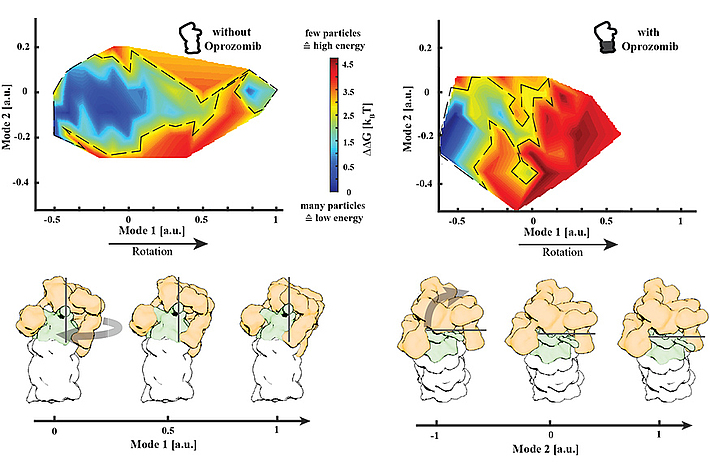
Our aim is to understand natural causes of malfunctioning macromolecular machines and the processes involved in the repair of malfunctioning machines on the example of the 26S proteasome.
Here especially, the unfolding and translocation of the substrate into the proteolytic chamber is prone to error. In extreme cases, the entrance to the 20S proteasome is clogged by a substrate and degradation can no longer take place. A few prominent examples of proteasome stalling are the surface antigen of the Eppstein-Barr virus or the amyloid-forming proteins huntingtin or ?-synuclein, which might directly clog the proteasome. Also, more general physiological situations such as oxidative stress, heat stress or arsenide intoxication can lead to a failure of the proteasome.
Due to its involvement in many processes, failure of the proteasome can lead to fatal consequences for the cell or even the organism. There are several rescue mechanisms known. In most cases, external protein factors bind to the periphery of the complex and change the degradation rate. However, in all cases is poorly understood how this is achieved mechanistically. Thus, we aim to use our biophysical method set to find the mechanisms that rescue stalled molecular machines.

